Building pharma's first industrial-grade digital twin engineWhere failures end in silico, success begins in reality.
Drug Development Pipeline
End-to-end modeling support
Discovery & Optimization
Preclinical
- Target identification and validation
- Model-supported lead optimization and candidate selection
- Best-in-class property identification and competitive benchmarking
Preclinical to Clinical
Translational
- Model-based characterization of therapeutic index
- Human efficacious dose and safe starting dose prediction
- Design of preclinical and phase I studies
Phase I-III Trials
Clinical
- Population PK/PD analysis
- Selection of RP2D and design of clinical phase II/III trials
- Patient stratification and biomarker selection
Scientific Disciplines
Core areas of expertise

Clinical Pharmacology
Support for Phases I, II, and III clinical trials for First-in-Human Dose (FHD), Single Ascending Dose (SAD), Multiple Ascending Dose (MAD), Drug-Drug Interaction (DDI), Bioavailability/Bioequivalence (BA/BE), and Food Effect.
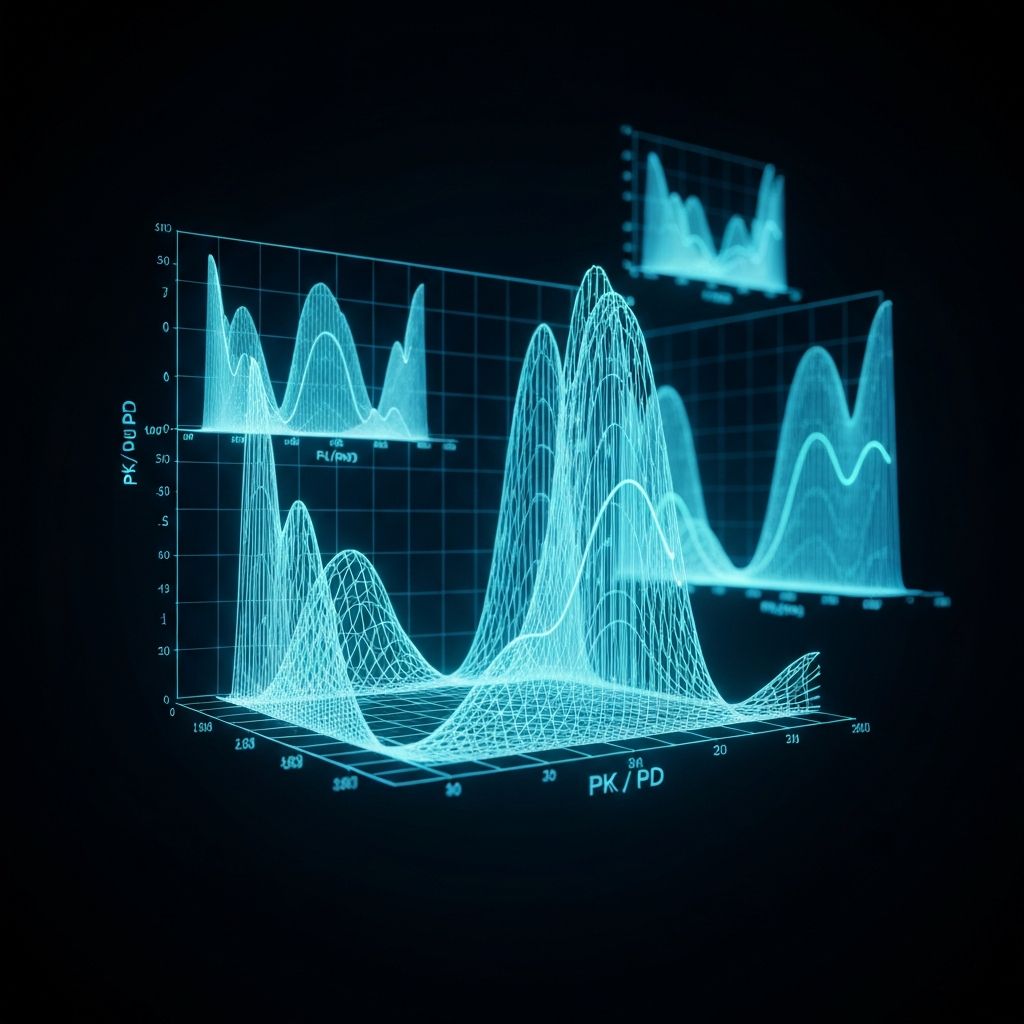
Pharmacometrics
Enhance drug development success with quantification of exposure-response to optimize clinical trials in time and resources, by preventing adverse events and providing the right dose to the right patient.
Paediatric expertise
We are experienced in providing full service consultative service crossing the research continuum in the following main areas.
Simulation Platforms
Proprietary AI-powered platforms
Antibody-Drug Conjugate
ADC Platform
Optimized design and dosing for reduced development risk

CAR-T & Gene Editing
Cell & Gene Platform
Precise modeling of delivery, expression, and pharmacodynamics
Dual-Target Binding
Bispecific Platform
Target-mediated disposition and therapeutic efficacy optimization

Systems Biology
Disease Modeling
Target efficacy evaluation and patient stratification
Get Started
Book a consultation
Connect with us to explore tailored strategies for navigating complex drug development challenges and accelerating the delivery of safe, effective therapies.